APPROXIMATION
Approximate 2D and 3D point set match
Approximate point set match is the task to align a search pattern represented by a set of 2D or 3D points in a, usually large, search space consisting of 2D or 3D points as well. The challenge is not to find only perfect matches, but allowing certain types of flexibility of the search pattern when embedded in the search space. The match itself might be only approximative, in other words, some deviation between the pattern and its counterpart in the search space is allowed. The measurement of the deviation can be based on the root mean square (RMSD) distance, but other distance functions (like average distance or maximum distance) might be of interest as well.
One example application is the search for known substructures of protein molecules in a database of proteins. However, approximate point set match has many other possible applications.
- We have implemented an application
 which finds substructures in proteins. Some results:
which finds substructures in proteins. Some results:
- The images show two locations where the 34 atoms of the active zone have been encountered in the protein 1IRU consisting of 47562 atoms. One is the perfect match (where the zone was cut) and one is an approximate match close by. The last image shows the optimal alignment.
IRU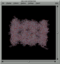 zona
zona  global
global  zoom
zoom  alignment
alignment 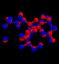
- The image shows the location of the best partial match (for the example the same zone as above was taken, but 6 atoms have been moved slightly).

- The image shows the five best locations where the five C-alpha atoms of the zone have been found in the protein.

- The images show two locations where the 34 atoms of the active zone have been encountered in the protein 1IRU consisting of 47562 atoms. One is the perfect match (where the zone was cut) and one is an approximate match close by. The last image shows the optimal alignment.
 in its server version is part of the algorithm suite of the Superimposé Web server for structural searches in protein data accessible at http://bioinformatics.charite.de/superimpose at the Charité Berlin, Germany.
in its server version is part of the algorithm suite of the Superimposé Web server for structural searches in protein data accessible at http://bioinformatics.charite.de/superimpose at the Charité Berlin, Germany.
Publications
-
Title: Superimposé: a 3D structural superposition server
Year: 2008
Journal: Nucleic Acids Research
Editorial: Oxford University Press
Issn: 0305-1048
Volumen: WSI
Pages: 47 - 54
Impact factor: 8.278 (Q1)
Base: JCR 2013
Area: Biochemistry and Molecular Biology
Type: Journal Articles
Authors: Arno Formella, Valentin Ziegler, Burghardt Wittig, Robert Preissner, Raphael .A Bauer, Thorsten Pöschel, P.E. Bourne, C. Frömmel, A. Goede, A. Guerler, A. Hoppe, E.W Knapp
-
Title: Búsqueda Aproximada de Patrones de Puntos en 2D y 3D
Year: 2007
Editorial: Fundación Alfredo Brañas [COLECCIÓN INFORMÁTICA 13/2007]
Isbn: 84-934497-0-9
Pages: 95 - 120
Type: Book Chapters
Authors: Arno Formella
People Involved
- Bauer, Raphael .A
- Formella, Arno
- González Fernández, Óscar
- Preissner, Robert
- Pöschel, Thorsten
- Sánchez Chao, Cástor
- Ziegler, Valentin
Student Work
-
Title: Búsqueda de patrones en cadenas de proteínas: Optimización de PSM
Year: 2009
Editorial: Universidade De Vigo [OUR PFC 2045]
Volumen: INX-886
Type: Bachelor's Thesis
Authors: Cástor Sánchez Chao
-
Title: PSM Server
Year: 2004
Volumen: INX-504
Type: Bachelor's Thesis
Authors: Óscar González Fernández
Press Articles
Resources
- Some interesting tools (v1.3) [Documents]


